Life Science and Technology News
Research on Lake Victoria Cichlids Uncovers the Processes of Rapid Species Adaptation
Scientists at Tokyo Institute of Technology (Tokyo Tech) have recently published the results of their investigations into adaptive radiation, which is when organisms rapidly evolve from an ancestral species into novel forms. Their genetic analyses of cichlids in Lake Victoria highlight several candidate genes that may drive adaptive radiation and provide evidence for decisive selective events that cause particular genetic variants to quickly gain dominance within populations. These findings broaden scientific understanding of how new species arise.
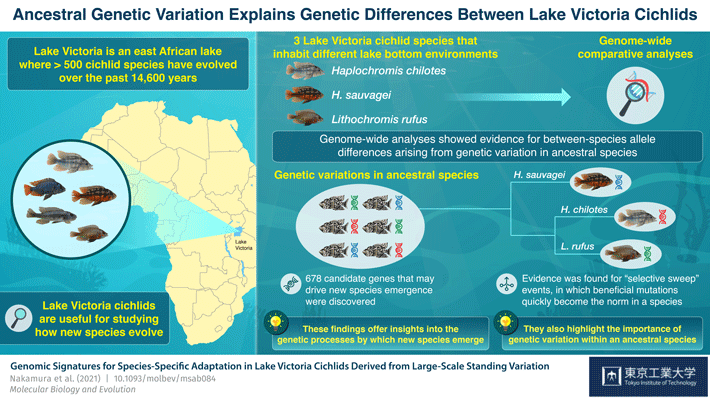
Biologists use the term adaptive radiation to describe a phenomenon in which new species rapidly evolve from an ancestral species, often in response to changes in the local environment that lead to new biological niches becoming available. To understand this process, biologists often turn to the cichlids of Lake Victoria, in which over 500 species of the fish have evolved over the past 14,600 years. As Professor Masato Nikaido of Tokyo Tech explains, "The level of genetic differentiation among species is considered very low due to the short period of time after these different species began evolving, and this limited genetic differentiation provides us with a great opportunity to find candidate genes that have contributed to adaptive radiation and the evolution of new species."
Past studies on Lake Victoria cichlids have shown that species living at different depths in the lake differ considerably with respect to the alleles for a gene that determines visual abilities. Spurred by this evidence, Professor Nikaido and his colleagues, including scientists from the Tanzania Fisheries Research Institute, wanted to learn more about the genetic processes underlying adaptation to different environmental conditions. They therefore analyzed the genomes of three species of Lake Victoria cichlids: Haplochromis chilotes and H. sauvagei, which inhabit only rocky shores, and Lithochromis rufus, which inhabits diverse lake-bottom environments, including rocky and sandy areas and areas with vegetation. Their findings are soon to appear in Molecular Biology and Evolution.![]()
Comparisons of genomic data from these different species led to the identification of 678 different genes located within genomic regions that exhibited considerable between-species variation. The researchers saw these genes as potentially playing critical roles in adaptive radiation, and they identified 43 that were specific to H. chilotes, 54 that were specific to H. sauvagei, and 63 that were specific to L. rufus. Some of these genes were related to behavioral variables, such as circadian rhythms, locomotion, and the development of sensory systems.
The genomic analyses also revealed signs of what biologists call "selective sweep events", in which selective pressures cause a beneficial mutation to rapidly increase its frequency within a population and become present in all members of a species. Furthermore, many of the species-specific alleles appeared to have existed alongside several alternative alleles prior to the adaptive radiation that produced different species of fish.
In conclusion, these analyses of Lake Victoria cichlids offer novel insights into the genetic processes underlying adaptive radiation. As Professor Nikaido puts it, "Our analyses uncovered the processes of species-specific adaption of Lake Victoria cichlids and the complexity of the genomic substrate that facilitated this adaptation."
The evidence for species-specific genetic alleles being derived from preexisting genetic variation is a particularly interesting finding that may prove useful to biologists who wish to understand adaptive radiation processes in different environments.
- Reference
| Authors : | Haruna Nakamura1, Mitsuto Aibara1, Rei Kajitani1, Hillary D.J. Mrosso2, Semvua I. Mzighani3,4, Atsushi Toyoda5, Takehiko Itoh1, Norihiro Okada6, and Masato Nikaido1 |
|---|---|
| Title of original paper : | Genomic Signatures for Species-Specific Adaptation in Lake Victoria Cichlids Derived from Large-Scale Standing Genetic Variation |
| Journal : | Molecular Biology and Evolution |
| DOI : | |
| Affiliations : | 1 School of Life Science and Technology, Tokyo Institute of Technology 2 Tanzania Fisheries Research Institute, Mwanza Fisheries Research Center 3 Tanzania Fisheries Research Institute, Headquarters 4 Tanzanian Fisheries Education and Training Agency 5 Comparative Genomics Laboratory, Japanese National Institute of Genetics 6 School of Pharmacy, Kitasato University |
| * Corresponding author's email: | E-mail : mnikaido@bio.titech.ac.jp |
- A pheromone-sensing gene that predates land-dwelling vertebrates| Life Science and Technology News
- Past genetic polymorphisms contributed to African cichlid diversity|Tokyo Tech News
- 【Labs spotlight】 Nikaido Laboratory|Life Science and Technology News
- Nikaido Lab. (Japanese)
- Researcher Profile|Tokyo Tech STAR Search - Masato Nikaido
- Researcher Profile|Tokyo Tech STAR Search - Mitsuto Aibara
- Researcher Profile|Tokyo Tech STAR Search - Takehiko Itoh
- Researcher Profile|Tokyo Tech STAR Search - Rei Kajitani
- Researcher Profile|Tokyo Tech STAR Search - Norihiro Okada
- Department of Life Science and Technology, School of Life Science and Technology
- National Institute of Genetics (NIG)
- Kitazato University
- Latest Research News
School of Life Science and Technology
—Unravel the Complex and Diverse Phenomena of Life—
Information on School of Life Science and Technology inaugurated in April 2016
Further Information
Associate Professor Masato Nikaido
School of Life Science and Technology, Tokyo Institute of Technology
E-mail : mnikaido@bio.titech.ac.jp