Life Science and Technology News
Working the Puzzle: Role of Sulfides in Aerobic/Anaerobic Switching in Bacteria
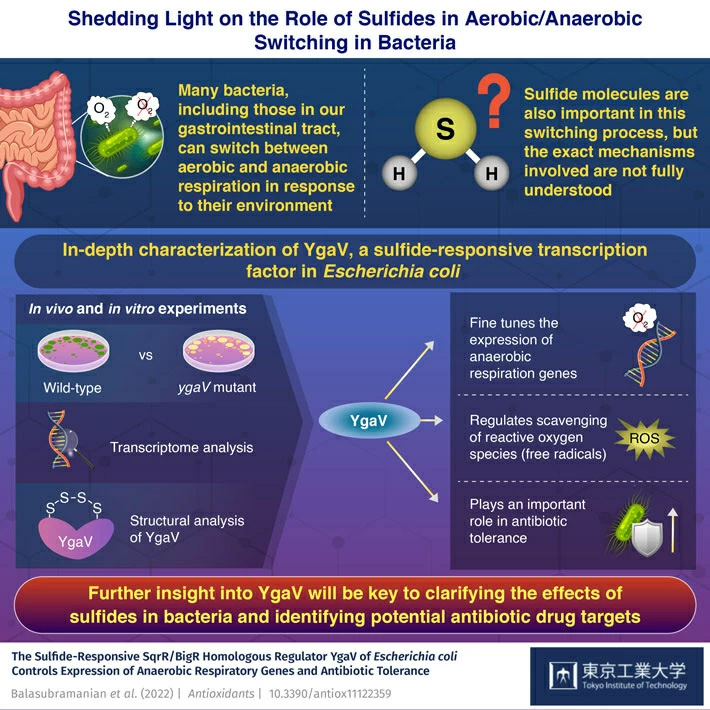
The YgaV protein found in the enteric bacterium Escherichia coli plays a critical role in maintaining homeostasis and antibiotic tolerance when exposed to sulfides, as shown in a recent study by researchers from Tokyo Tech. Comparisons between normal and ygaV mutant bacterial strains shed light on the many effects of the YgaV transcription factor. This will lead to a better understanding of aerobic/anaerobic respiration switching and the link between antibiotic tolerance and free radicals.
The bacteria in our intestinal tract play many essential roles in maintaining our health and metabolism. Keeping these organisms happy and striking a good balance among species is therefore very important to remain healthy. Many of these bacteria have special mechanisms by which they can switch from aerobic respiration (using oxygen) to anaerobic respiration (without using oxygen) depending on their environment.
This peculiar switching process is also involved in other functions, such as increasing antibiotic tolerance and protecting the cell against reactive oxygen species (ROS), a family of potentially harmful free radicals. Interestingly, small sulphur-containing molecules called sulfides also affect this switching process. Bacteria seem to have evolved ways to sense hydrogen sulfide (H2S) levels and adapt their metabolism accordingly. How exactly they do this, however, is not well understood.
To address this knowledge gap, a research team including Associate Professor Shinji Masuda from Tokyo Institute of Technology, Japan, recently focused on YgaV, a transcription factor found in the enteric bacterium Escherichia coli that could add some pieces to the puzzle. In their latest paper, which was published in Antioxidants![]() , the team explains that YgaV is evolutionarily related (and almost identical) to certain proteins previously discovered in other bacteria that respond to H2S levels. This motivated them to decipher what the role of YgaV is and how it helps E. coli thrive when the chemical conditions around them change.
, the team explains that YgaV is evolutionarily related (and almost identical) to certain proteins previously discovered in other bacteria that respond to H2S levels. This motivated them to decipher what the role of YgaV is and how it helps E. coli thrive when the chemical conditions around them change.
The researchers conducted a series of experiments in normal (or WT, for 'wild-type') and mutant E. coli strains, the latter of which had a non-functional ygaV gene. They exposed these cultures to different atmospheric conditions (oxygen-rich, oxygen-deficient, and H2S-rich). Then, through transcriptome analysis, the team investigated which genes were expressed differently between the WT and mutant strains, thus highlighting the role of YgaV. They also exposed both strains of bacteria to several types of antibiotics and compared their effects. Further, they measured the cellular concentration of oxygen peroxide (H2O2), a type of ROS, and analyzed some of the structural details of how YgaV binds to sulphur atoms.
Overall, the results showed that YgaV is crucial for fine-tuning the expression of various anaerobic respiratory genes and managing the levels of ROS in response to external H2S. Notably, the antibiotic tolerance was different between strains, as Associate Professor Masuda remarks: "While the WT bacteria showed increased antibiotic tolerance under H2S-atmospheric conditions, the ygaV mutant did not. Additionally, antibiotic sensitivity was higher in the mutant than in the WT strain, both in the presence and absence of external H2S."
These findings clarify some of the mechanisms by which various bacteria can regulate their internal chemical balance in response to changes in their surroundings. This knowledge could be used to identify potential drug targets to make pathogenic bacteria more susceptible to antibiotics. In turn, this would help in the treatment of infectious diseases. Most importantly, the link that exists between YgaV, antibiotic tolerance, and ROS levels can be studied in more detail to gain a better understanding of how antibiotic resistance works. "Further research on YgaV may contribute to solving the decade-old enigma of whether there is a relationship between ROS generation and antibiotic tolerance," notes Associate Professor Masuda.
There is still much more to learn about these microscopic organisms, but we are certainly getting there one step at a time!
- Reference
| Authors : | Rajalakshmi Balasubramanian1, Koichi Hori1, Takayuki Shimizu2, Shingo Kasamatsu3, Kae Okamura3, Kan Tanaka4, Hideshi Ihara3, and Shinji Masuda1,* |
|---|---|
| Title : | The Sulfide-Responsive SqrR/BigR Homologous Regulator YgaV of Escherichia coli Controls Expression of Anaerobic Respiratory Genes and Antibiotic Tolerance |
| Journal : | Antioxidants |
| DOI : | |
| Affiliations : | 1 Department of Life Science and Technology, Tokyo Institute of Technology 2 Graduate School of Arts and Sciences, The University of Tokyo 3 Department of Biological Chemistry, Graduate School of Science, Osaka Metropolitan University 4 Laboratory for Chemistry and Life Science, Institute of Innovative Research, Tokyo Institute of Technology |
| * Corresponding author's email: | shmasuda@bio.titech.ac.jp |
- Sulfide-sensing mechanisms in purple bacteria│Life Science and Technology News
- Cultivating a new crop of superplants--a lesson from the fittest│Tokyo Tech News
- Masuda Lab.
- Shinji Masuda | Researcher Finder - Tokyo Tech STAR Search
- Koichi Hori | Researcher Finder - Tokyo Tech STAR Search
- Department of Life Science and Technology, School of Life Science and Technology
- The University of Tokyo
- Osaka Metropolitan University
- Latest Research News
School of Life Science and Technology
—Unravel the Complex and Diverse Phenomena of Life—
Information on School of Life Science and Technology inaugurated in April 2016
Further Information
Associate Professor Shinji Masuda
School of Life Science and Technology, Tokyo Institute of Technology
E-mail : shmasuda@bio.titech.ac.jp