Life Science and Technology News
Sharing Shears: Conserved Protein Segment Activates Molecular DNA Scissors for DNA repair
Scientists at Tokyo Institute of Technology (Tokyo Tech) have uncovered mechanisms underlying the activation of the MRN complex- the cell's DNA scissors. Using purified yeast proteins, they demonstrated that phosphorylation of Ctp1, a homolog of a tumor-suppressor protein, plays a key role in activating MRN complex's DNA clipping activity. Intriguingly, a short segment of yeast Ctp1 or its human counterpart could stimulate endonuclease activity of their respective MRN complexes, suggesting its conserved function across species.
DNA functions as a roadmap that guides the identity and functions of cells. A glitch in the DNA can have serious deleterious effects resulting in the malfunction or loss of crucial proteins, thus affecting normal cellular function and viability. These glitches often manifest as double stranded breaks in the DNA that may occur spontaneously or from exposure to certain chemicals. To deal with these kinks, cells have evolved a DNA repair machinery that scans, identifies, and fixes breaks in the DNA by ligating the gaps. However, DNA breaks often have "dirty ends" that cannot be directly ligated or sealed as they are unexposed or blocked by certain proteins or irregular chemical structures. Such DNA ends thus need to first be clipped and freed so that it can be processed further. Moreover, such end-resection of DNA break ends are prerequisite for them to be accurately repaired by homologous recombination. Among such molecular scissors, or nuclease enzymes, Mre11 is a key player.
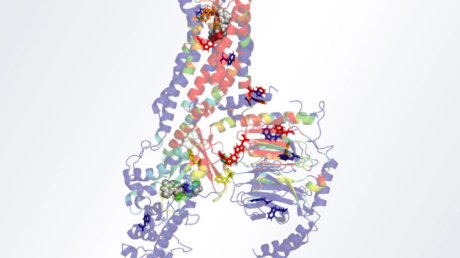
Mre11 teams up with proteins Rad50 and Nbs1 to collectively form the ‘MRN’ complex. The interaction of this complex with the tumor-suppressor protein CtIP in humans, has been shown to trigger the DNA clipping function of the complex (Figure 1). However, the mechanisms underlying this interaction have hitherto remained unexplored.
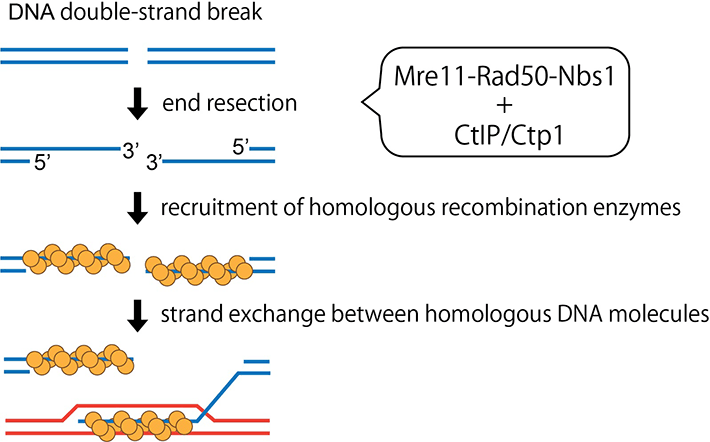
Figure 1. MRN complex stimulating activity of CtIP/Ctp1
CtIP/Ctp1 stimulates endonuclease activity of the MRN complex, which not only polishes "dirty" ends of DNA breaks, but promotes accurate repair of DNA breaks through the homologous recombination mechanism.
Now, Assistant Professor Hideo Tsubouchi and Professor Hiroshi Iwasaki from Tokyo Institute of Technology and their team have decoded the stepwise interaction and activation of the MRN complex using Ctp1 proteins in yeast, which are homologous to the human CtIP. Discussing their findings that have recently been published in PNAS![]() , Iwasaki says, "The MRN complex is pivotal in the homologous recombination-mediated repair of DNA double stranded breaks. To better understand how CtIP influences the activity of the MRN complex, we purified yeast proteins and quantified their interactions."
, Iwasaki says, "The MRN complex is pivotal in the homologous recombination-mediated repair of DNA double stranded breaks. To better understand how CtIP influences the activity of the MRN complex, we purified yeast proteins and quantified their interactions."
The scientists found that phosphorylation or the addition of phosphate groups to Ctp1 was the key first step in activating the MRN complex. More specifically, phosphorylation enabled the physical interaction of Ctp1 with the Nbs1 protein of the complex, which was vital for subsequent endonuclease stimulation. The DNA clipping activity was extremely poor when the MRN complex was mixed with unphosphorylated Ctp1.
Furthermore, the scientists identified a short stretch of merely 15 amino acids at the C-terminal region of Ctp1 that was indispensable for the endonuclease activity of the Ctp1-stimulated MRN. Moreover, a synthetic peptide mimicking this region of Ctp1 or CtIP was able to activate the yeast or human MRN complex, respectively, suggesting that the function of the C-terminal Ctp1 is likely conserved across species and is the ultimate determinant in MRN activation.
Excited about the prospective application of their findings, Tsubouchi remarks, "Modification of the CT15 peptide can yield a strong activator or potential inhibitor of the MRN complex. Targeting this endonuclease activity can have potentially useful applications in homologous recombination-based gene editing."
With rapid advancements in recombinant DNA and molecular medicine, these findings could empower geneticists to unravel the mysteries of the genome and identify the hidden intricacies of genetic disorders with greater ease and effectiveness in the days to come.
- Reference
| Authors : | Aleksandar Zdravkovic, James M. Daley, Arijit Dutta, Tatsuya Niwa, Yasuto Murayama, Shuji Kanamaru, Kenatro Ito, Takahisa Maki, Bilge Argunhan, Masayuki Takahashi, Hideo Tsubouchi*, Patrick Sung* and Hiroshi Iwasaki* |
|---|---|
| Title of original paper : | A conserved Ctp1/CtIP C-terminal peptide stimulates Mre11 endonuclease activity |
| Journal : | PNAS (2021) |
| DOI : |
10.1073/pnas.2016287118
|
| Affiliations : | Graduate School of Bioscience and Biotechnology, Tokyo Institute of Technology
Institute of Innovative Research, Tokyo Institute of Technology
Department of Biochemistry and Structural Biology, University of Texas Health Science Centre
Centre for Frontier Research, National Institute of Genetics |
| * Corresponding author's email: | htsubouchi@bio.titech.ac.jp, hiwasaki@bio.titech.ac.jp |
- Helping a Helper: Uncovering How Different Proteins Cooperate in DNA Repair | Life Science and Technology News
- Scientists unraveling the molecular details of DNA recombination regulation | Life Science and Technology News
- Elucidation of the Complicated Regulating Mechanism in Homologous Recombination |Tokyo Tech News
- 【Labs spotlight】 Iwasaki Laboratory | Life Science and Technology News
- Iwasaki Laboratory
- Researcher Profile | Tokyo Tech STAR Search - Aleksandar Zdravkovic
- Researcher Profile | Tokyo Tech STAR Search - Hideo Tsubouchi
- Researcher Profile | Tokyo Tech STAR Search - Hiroshi Iwasaki
- Cell Biology Center, Institute of Innovative Research
- Institute of Innovative Research (IIR)
- Department of Life Science and Technology, School of Life Science and Technology
- The Graduate School of Biomedical Sciences at UT Health San Antonio
- Latest Research News
Contact
Public Relations Group, Tokyo Institute of Technology
E-mail : media@jim.titech.ac.jp
Tel +81-3-5734-2975