Life Science and Technology News
【Labs spotlight】 Takinoue Laboratory

Areas of Supervision
・Major, Department
Primary/ Artificial Intelligence, Computer Science
Secondary/Life Science and Technology, Life Science and Technology/ Others
Professor Masahiro TAKINOUE![]()
| Office | Room 1806, J2 building, Suzukakedai campus |
|---|---|
| Degree | Ph.D.2007, Department of Physics, The University of Tokyo |
| Areas of Research | Biophysics, DNA Nanotechnology, Synthetic Biology, Soft Matter Physics, Molecular Robotics, Microfluidcs |
| Keywords | Artificial cells, Artificial nuclei, DNA gel, DNA origami, Microdroplet |
| Web site | Takinoue Lab. |
Research interest
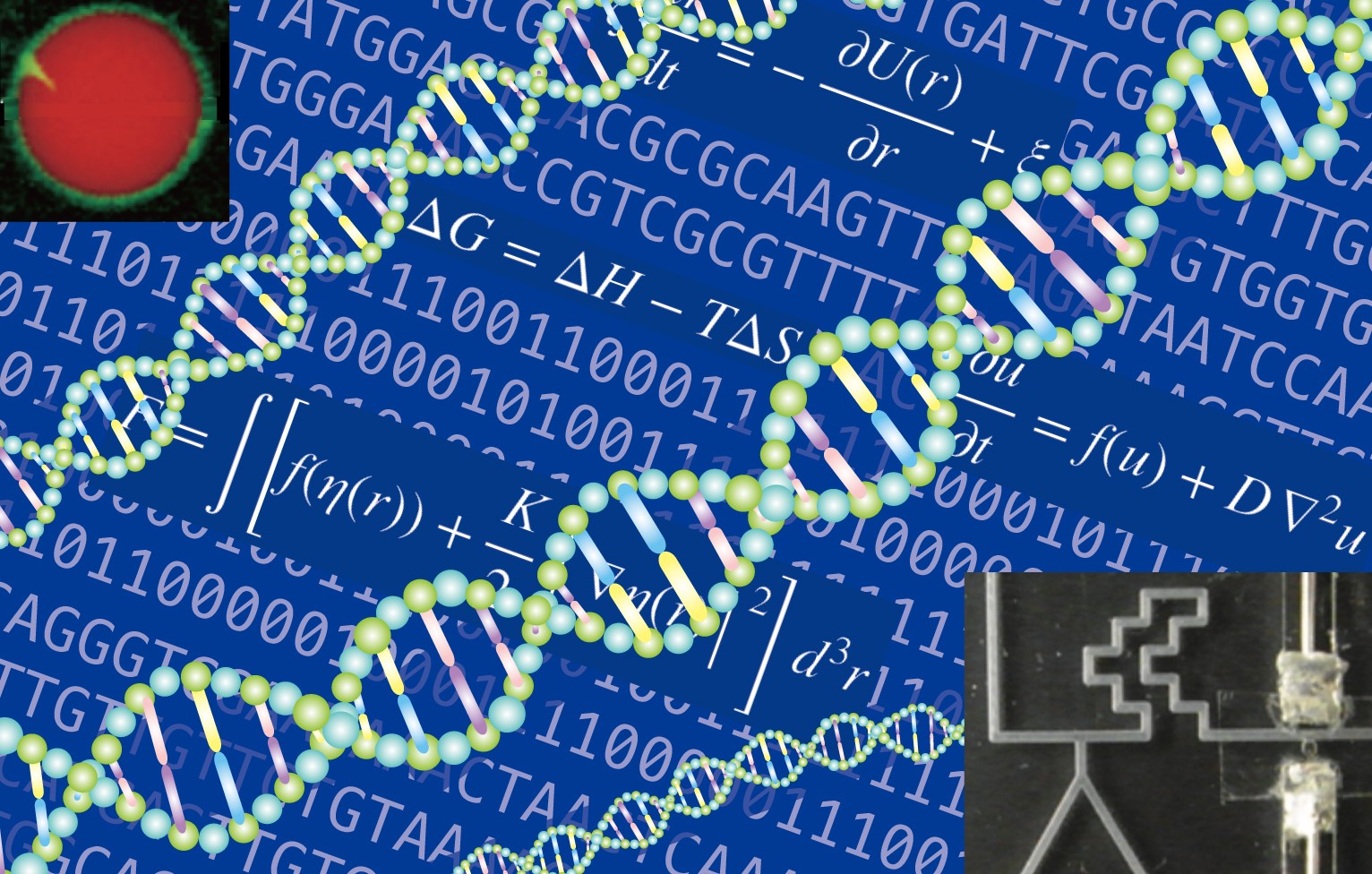

The life system is one of the most complex systems in nature. We can see dynamical autonomous behaviors, emergence of functional behaviors, and also intelligence in the life systems unlike non-living systems. Although the Life systems are complex, the life systems are also composed of usual matter same as non-living systems. The research themes of our lab are basically inspired by such physical point of view of life systems.
Our group studies the construction of bio-inspired dynamical self-organized systems (molecular robots, artificial cells) using information molecules such as DNA. Through their construction, we are challenging to understand "What is Life?" from a physical point of view about nano/micro-scale autonomous systems and to invent novel post-silicon computing-based intelligent molecular machines, molecular computers, and molecular robots with sophisticated functions beyond life systems.
(I) Molecular Robotics and Molecular Computing by nanobiotechnology
We are challenging to construct autonomous nano/micrometer-sized "molecular robots" and "molecular computers". Molecular robots and molecular computers are made of biopolymers and biomaterials such as DNA, protein, lipids, etc. The Molecular robots and molecular computers have autonomously working sensors, processors, and actuators. They can work in microscopic spaces like inside of cells and can be applied to medical systems, tiny machines, etc.
(II) Artificial Cell and Nonlinear Non-equilibrium Science with Microfluidics
We are constructing "artificial cells" using biomolecules (DNA, RNA, protein, lipid, etc.) to understand essential aspect of life systems. Life systems exhibit dynamical behavior such as autonomous sensing, autonomous information processing, autonomous motion, etc. Artificially constructing cell-like systems is important not only for basic science (physics, chemistry, biology) but also for applied science and engineering.
The recent progress is as follows
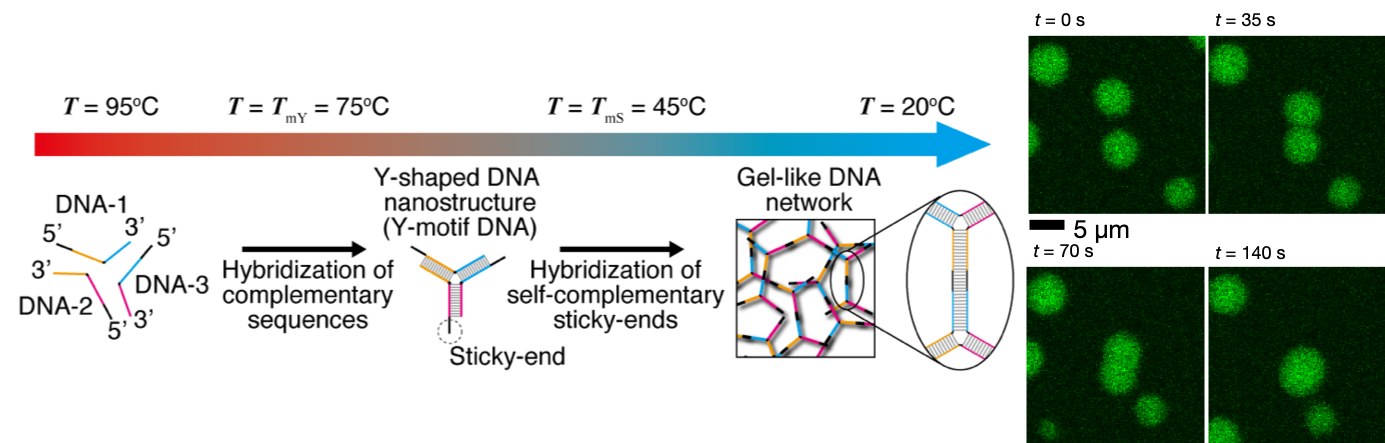
(1) Construction of DNA microparticles/droplet based on phase transition and phase separation of DNA gels
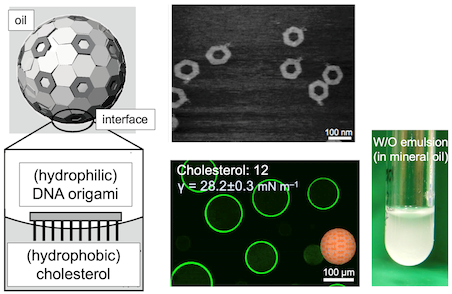
(2)Artificial cells based on DNA origami nanoplate (Ref: [1])
(3)DNA cytoskeleton for stabilizing liposome-based artificial cells (Ref: [2])
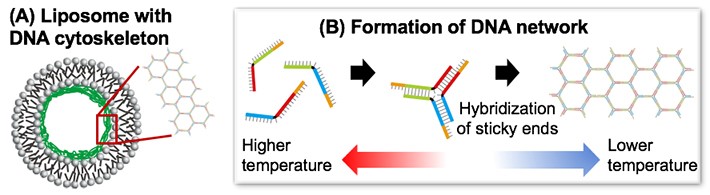
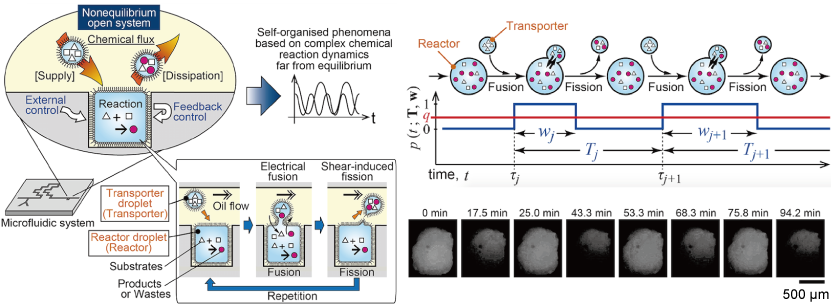
(4)Non-equilibrium open microreactor based on droplet artificial cell (Ref: [3])
Selected publications
Original articles
- [1]D. Ishikawa, Y. Suzuki, C. Kurokawa, M. Ohara, M. Tsuchiya, M. Morita, M. Yanagisawa, M. Endo, R. Kawano, *M. Takinoue, "DNA Origami Nanoplate-Based Emulsion with Designed Nanopore Function", ChemRxiv, Preprint. DOI: 10.26434/chemrxiv.7046495.v1 link
- [2]C. Kurokawa, K. Fujiwara, M. Morita, I. Kawamata, Y. Kawagishi, A. Sakai, Y. Murayama, S.-i. M. Nomura, S. Murata, *M. Takinoue, *M. Yanagisawa, "DNA cytoskeleton for stabilizing artificial cells", Proc. Natl. Acad. Sci. USA, 114(28), 7228-7233, (2017). doi: 10.1073/pnas.1702208114. link Press release: [Japanese] [English]
- [3]H. Sugiura, M. Ito, T. Okuaki, Y. Mori, H. Kitahata, *M. Takinoue, "Pulse-density modulation control of chemical oscillation far from equilibrium in a droplet open-reactor system", Nature Commun., 7, 10212, (2016). DOI: 10.1038/ncomms10212 link Press release: [Japanese] [English]
Review articles
- [4]*Y. Sato, *M. Takinoue, "Creation of artificial cell-like structures promoted by microfluidics technologies", Micromachines, Vol.10(4), 216, (2019), DOI: 10.3390/mi10040216
- [5]M. Takinoue, *S. Takeuchi, "Droplet microfluidics for the study of artificial cells", Anal. Bioanal. Chem., vol. 400, pp. 1705-1716 (2011). DOI: 10.1007/s00216-011-4984-5

*Find more about the lab and the latest activities at Takinoue Lab.![]()
*May 1, 2025:Some of the content has been updated with the latest information.