Life Science and Technology News
【Labs spotlight】Uriu Laboratory
The Department has a variety of laboratories for Life Science and Technology, in which cutting-edge innovative research is being undertaken not only in basic science and engineering but also in the areas of medicine, pharmacy, agriculture, and multidisciplinary sciences. This "Spotlight" series features a laboratory from the Department and introduces you to the laboratory's research projects and outcomes. This time we focus on Uriu Laboratory, studying developmental biology and chronobiology using mathematical modeling and simulations.

Areas of Supervision
Primary/Life Science and Technology
Secondary/Science and Technology for health Care and Medicine, Human Centered Science and Biomedical Engineering
Associate Professor Koichiro Uriu![]()
| Office | Room 302A, Midorigaoka building 6, Ookayama campus |
|---|---|
| Degree | PhD 2010, Kyushu University |
| Areas of Research | Mathematical Biology, Developmental Biology, Chronobiology |
| Keywords | Simulation, Somitogenesis, Circadian clock |
Research interest
Mathematical Biology
Biology is full of “numbers”, such as the number of animals in Ecology and the concentration of proteins in cells in Cell Biology. Mathematics is the research area that addresses the “numbers”. Therefore, it is quite reasonable to study Biology from a mathematical perspective. In recent years, mathematical models have been getting attention and used in various fields of Biology. Mathematical models can help us understand the role of complex gene regulation in cells. Models can also be used to simulate the spread of infectious diseases. Importantly, abstraction of biological phenomena in mathematical equations will allow us to reveal universal laws that hold beyond species. The field that uses mathematics in life sciences is called "Mathematical Biology”. In our laboratory, we use mathematical approaches to study (1) segment formation observed in vertebrate development and (2) mammalian circadian clocks.
Body segment formation in vertebrates
Vertebrate body includes segmental structures such as vertebrae and ribs. These segmental structures are derived from mesodermal tissues called somites during development. Somites are formed one by one from the head to the tail in a species-specific temporal period (5 hours for human, 2 hours for mouse, 1.5 hours for chick, and 0.5 hours for zebrafish). It has been known that the period of segment formation is determined by the oscillation of protein concentrations in cells. This oscillation acts as a "clock" that determines the timing of somite formation, and is known as a "segmentation clock”. Cells communicate with neighboring cells to synchronize the segmentation clock (protein rhythms) to differentiate into a somite at the same time. In our laboratory, we use mathematical models to study the mechanism of synchronization of segmentation clock, and the role of tissue elongation and cell movement on segment formation.
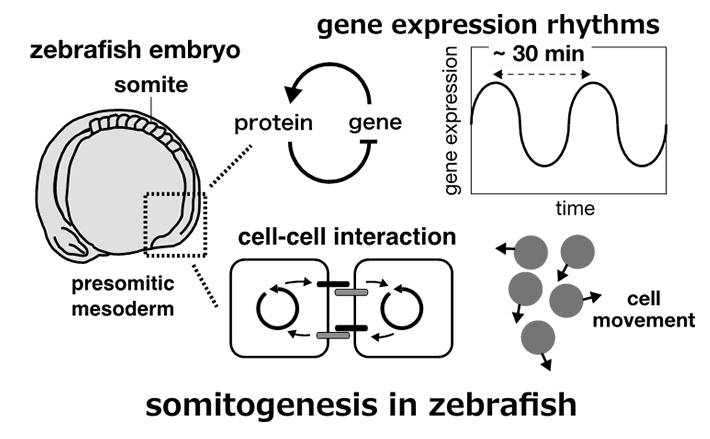
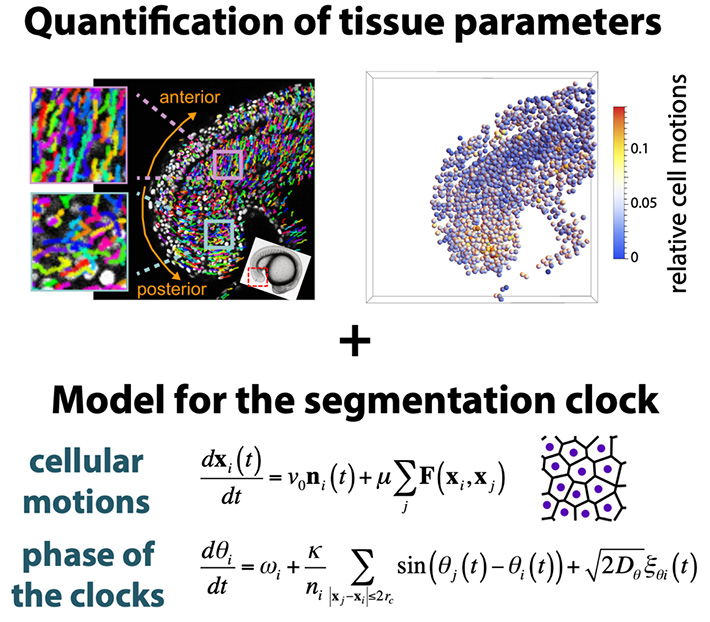
Cell movement and synchronization of clocks: somites bud off from the anterior part of a unsegmented tissue, the presomitic mesoderm. Each cell can communicate with neighboring cells with membrane proteins, Delta ligands and Notch receptors. In addition, cells in the presomitic mesoderm move around in the tissue, exchanging neighboring cells over time. Here is a question: how do cells synchronize segmentation clock if they move and frequently exchange their neighbors to communicate? It would be nice if one can study the effects of cell movement on synchronization by experimental approaches with living animals, but such experiments are currently difficult. To answer this question, we choose a theoretical approach; we quantified cell movement by tracking cells in imaging data, and simulated cell movement on a computer to see how movement affects synchronization of the clock. We found that cell movement and neighbor exchange promote synchronization of the segmentation clock. The synchronization with neighboring cells is indeed disrupted by movement. However, movement also creates a chance to meet and communicates with cells that were previously distant from each other. Therefore, at a cell population level, cell movement facilitates information transfer, realizing quick synchronization. Our study suggests that cell movement makes the segmentation clock more precise and allows for robust somite formation in vertebrate development.
Mammalian Circadian Clock
Almost all living organisms, including humans, have an internal clock that measures 24-hour cycles, known as the circadian clock. The circadian clocks regulate behavioral and physiological rhythms with nearly 24-hour period in these organisms. In humans, the pacemaker of the circadian clock is located in the brain tissue suprachiasmatic nucleus (SCN), which consists of tens of thousands of neurons. In these neurons, expression of circadian clock genes oscillates in approximately 24-hour period. In mammals, several circadian clock genes such as Period (Per), Cryptochrome (Cry), Bmal1, and Clock have been identified. Regulatory network of these circadian clock genes is the basis for the generation of transcriptional rhythms. For example, in mammals, the CLOCK-BMAL1 complex induces transcription of Per and Cry. Translated PER and CRY protein then form complex and repress the transcriptional activity of CLOCK-BMAL1. This negative feedback regulation by Per and Cry is essential for generating circadian rhythms. In our laboratory, we study the mechanisms for causing gene expression rhythms, the response of the circadian clock to light stimuli and compound administration, and the temperature compensation of the circadian period.
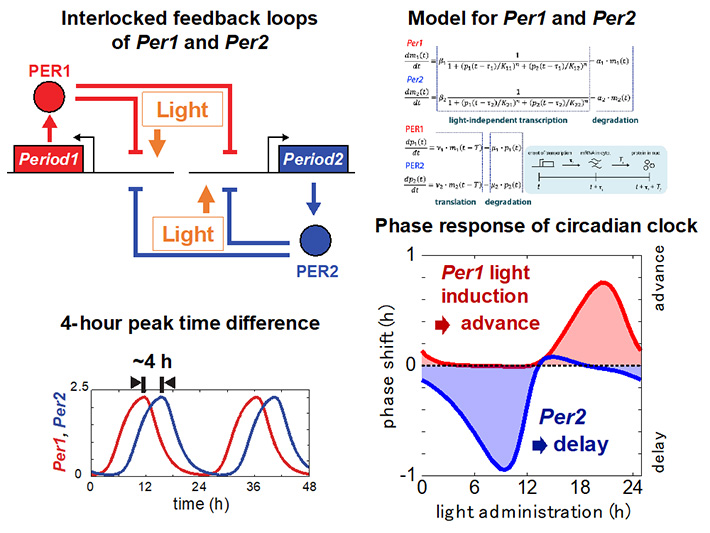
Phase response of the circadian clock: Light input from the retina is transmitted to the SCN and alters the gene expression rhythms in SCN neurons. Depending on the time of day when the light stimulus is received, the expression rhythm of circadian clock gene is either advanced or delayed. This allows the circadian clock to entrain to the light-dark cycle, so that, for example, when we travel abroad, we can adjust our circadian clock to the local time within a few days. Light information is input to two circadian clock genes, Per1 and Per2. Because of the homology of amino acid sequences between PER1 and PER2 proteins, we would expect that these two proteins have similar functions. Are Per1 and Per2 simply two redundant circadian clock genes with the same function? There is a phase difference between the expression rhythms of Per1 and Per2, with Per2 expression peaking about 4 hours after the peak of Per1 transcription. We have shown by mathematical modeling that this 4-hour time difference results in distinct roles in the light response; Per1 advances the circadian clock upon light stimulation, while Per2 delays it. These results suggest that if, for example, a compound that specifically induces the transcription of only Per1 is identified, the circadian clock can be advanced independently of administration timing. Such a compound would have potential as a therapeutic agent for jet lag and sleep disorders. Our laboratory is collaborating with an experimental laboratory to validate these theoretical predictions.
Selected publications
Taijiro Yabe, Koichiro Uriu, Shinji Takada. (2023) Ripply suppresses Tbx6 to induce dynamic-to-static conversion in somite segmentation. Nature communications 14(1) 2115-2115
Koichiro Uriu, Hajime Tei. (2021) Complementary phase responses via functional differentiation of dual negative feedback loops. PLOS Computational Biology 17(3) e1008774
Koichiro Uriu, Bo-Kai Liao, Andrew C Oates, Luis G Morelli (2021) From local resynchronization to global pattern recovery in the zebrafish segmentation clock. eLife 10
Koichiro Uriu, Hajime Tei (2019) A saturated reaction in repressor synthesis creates a daytime dead zone in circadian clocks. PLOS Computational Biology 15(2) e1006787
Koichiro Uriu, Rajasekaran Bhavna, Andrew C. Oates, Luis G. Morelli (2017) A framework for quantification and physical modeling of cell mixing applied to oscillator synchronization in vertebrate somitogenesis. Biology Open 6(8) 1235-1244
Koichiro Uriu, Luis G. Morelli (2014) Collective cell movement promotes synchronization of coupled genetic oscillators. Biophys. J. 107(2) 514-526
Koichiro Uriu, Saul Ares, Andrew C. Oates, Luis G. Morelli (2013) Dynamics of mobile coupled phase oscillators. Phys. Rev E 87(3)
Luis G. Morelli, Koichiro Uriu, Saul Ares, Andrew C. Oates (2012) Computational approaches to developmental patterning. Science 336(6078) 187-191
Koichiro Uriu, Yoshihiro Morishita, Yoh Iwasa (2010) Random cell movement promotes synchronization of the segmentation clock. Proc. Nat. Acad. Sci. USA 107(11) 4979-4984
Contact
Associate Professor Koichiro Uriu
Room 302A, Midorigaoka building 6, Ookayama campus
E-mail : uriu@bio.titech.ac.jp
Tel / Fax : +813-5734-3582
*May 1, 2025:Some of the content has been updated with the latest information.