Life Science and Technology News
Protein intentionally terminates own synthesis by destabilizing synthesis machinery - the ribosome
A joint research group of Tokyo Institute of Technology (Tokyo Tech) and Kyoto Sangyo University has discovered that a protein, during its synthesis, may destabilize the structure of the ribosome1 and end its own synthesis prematurely, and found that this phenomenon is used for adapting the cell to its environment.
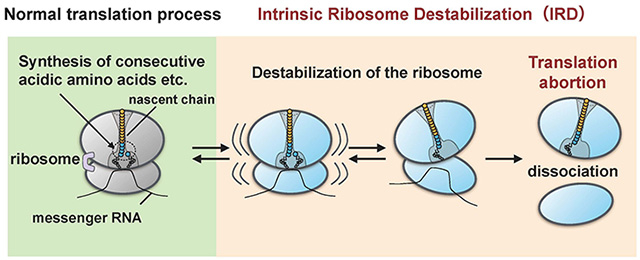
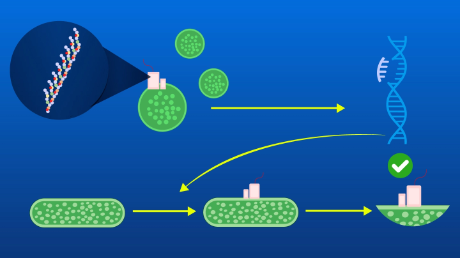
Figure 1. Destabilization and dissociation of the ribosome
The ribosome becomes destabilized by the protein (nascent chain) it itself synthesized (IRD) and splits apart into the large and small subunits. As a result, the protein synthesis ends mid-cycle without reaching the end point.
Background
Proteins are functional molecules, which support life and are three-dimensional structures of polypeptide chains, chains of linked amino acids. The configuration of amino acids on this chain (the sequence) is written in the DNA sequence. Polypeptide chains are formed where the ribosome, the protein synthesis machinery, resides on a copy of the DNA sequence (messenger RNA). It starts polymerizing amino acids one by one by selecting from 20 types of amino acids to add to the link according to the genetic code. It is completed at the endpoint where the chain detaches from the ribosome.
The process of a protein being synthesized at a ribosome is called "translation", and all proteins of all living organisms, including humans, are produced through translation. It had been thought that, in the process of linking amino acids at the ribosome, the newly formed polypeptide chain (nascent chain) was synthesized continuously.
In recent studies by this research group and others, it has become apparent that fluctuation in speed occurs significantly in translation—that some nascent chains, depending on the amino acid sequence, act on the ribosome that is forming it to slow down the translation steptime.
Overview of the Research Achievement
The research group led by Hideki Taguchi at Tokyo Tech and Koreaki Ito at Kyoto Sangyo University found that, when a reconstituted cell-free translation system2 of E. coli was made to translate proteins having sequences of about ten acidic amino acids (aspartic and glutamic acids) or sequences in which either acidic amino acid and an amino acid called proline are linked alternately, translation stops midway when such a sequence has been translated. This abortive event occurs when the ribosomes synthesizing the amino acid sequence become destabilized as a result of being acted on by the nascent chain (the researchers designate this "IRD": "intrinsic ribosome destabilization"). It results in the ribosome splitting apart into the large and small subunits (Figure 1). The ribosome, with its role of synthesizing several thousands or tens of thousands of types of proteins within the cell, had been thought to be in command in linking any combinations of amino acid sequences. However, the discovery of IRD, in which the ribosome is destabilized during translation by nascent chains it itself is synthesizing, indicates that translation is something which progresses with the potential risk of being aborted. Proteins which include amino acid sequences that trigger IRD are not able to complete their syntheses.
At first glance, this phenomenon of IRD may seem like a defect of the ribosome, but what does it indicate for living organisms?
The researchers had discovered that living organisms also possess a mechanism to counteract IRD. This finding led them to use a mutant of E. coli missing this mechanism, with ribosomes thus slightly destabilized and prone to IRD, and analyze all the protein inside a cell (the proteome). As a result, the team found that, in the mutant, the cellular amounts of a number of proteins varied when compared to the wild strain. In particular, MgtA, a membrane protein which transports magnesium ion into cells, was found to be expressed over ten times more. Interestingly, the gene called mgtL and regulating the expression of MgtA possessed an IRD sequence. The result of the analysis showed that E. coli regulates the expression of MgtA, using a special mechanism featuring the IRD phenomenon due to the sequence of mgtL, in response to changes in the magnesium concentrations in the cell.
Magnesium ions are necessary for many life processes inside the cell, and is required for translation, in particular, to stabilize the ribosome. Thus, the findings of this research suggest that E. coli acquired a mechanism for maintaining the cellular magnesium ion concentration by using mgtL's IRD ability to express an abundance of MgtA when its growth environment becomes poor in magnesium. In other words, a living organism has a mechanism for monitoring changes in the intracellular environment by taking advantage of the phenomenon of IRD (Figure 2).
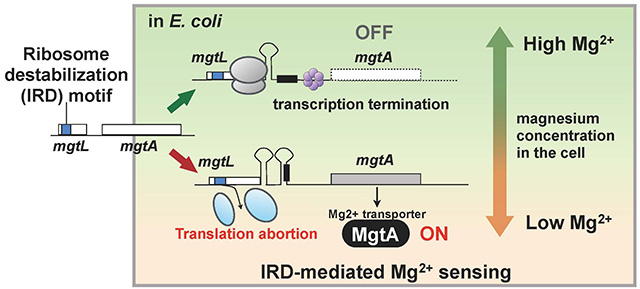
Figure 2. Mechanism for sensing magnesium ions resulting from ribosome destabilization
The gene, mgtL, controlling the expression of the membrane protein MgtA which transports magnesium into cells has a ribosome-destabilizing sequence, and translation is aborted when the magnesium concentration inside the cell is low. There is a mechanism which turns on MgtA synthesis when mgtL translation is aborted, and it is thought that the resulting abundant expression of MgtA raises the magnesium ion concentration inside the cell.
Future Development
This study revealed that, in addition to containing information that determines three-dimensional structures of proteins, amino acid sequences even hold the ability to influence the stability of the machinery synthesizing it and to abort translation mid-cycle, eventually. This furthers our comprehension of the foundation of life processes, which are supported by regulated expression of genetic information. Early, premature cessation of translation had not been considered to be a vital process in the life sciences to date. Taguchi and the team members believe that their accomplishment can be expanded, in conjunction with the current prosperity of life sciences, to various applications such as the production of new useful proteins or the development of biopharmaceuticals.
a large composite made of RNA and protein, which synthesizes protein constituents (polypeptide chains). It reads the nucleotide sequence of messenger RNAs and selects from the 20 types of amino acids according to the genetic code on the message, then connects them in the order specified.
2 Reconstituted cell-free translation system
an in vitro protein synthesis system containing only the purified components necessary for synthesizing proteins.
Reference
| Authors : | Yuhei Chadani1, Tatsuya Niwa1, Takashi Izumi1, Nobuyuki Sugata1, Asuteka Nagao1, Tsutomu Suzuki2, Shinobu Chiba3, Koreaki Ito3,*, Hideki Taguchi1,* |
|---|---|
| Title of original paper : | Intrinsic Ribosome Destabilization Underlies Translation and Provides an Organism with a Strategy of Environmental Sensing |
| Journal : | Molecular Cell 68, 528-539 (2017) |
| DOI : | 10.1016/j.molcel.2017.10.020 |
| Affiliations : |
1Cell Biology Center, Institute of Innovative Research, Tokyo Institute of Technology
2Department of Chemistry and Biotechnology, Graduate School of Engineering, The University of Tokyo 3Faculty of Life Sciences and Institute for Protein Dynamics, Kyoto Sangyo University |
- Taguchi Laboratory
- Hideki Taguchi - Chaperones supporting protein life cycles| Research Story
- Labs spotlight #41 - Taguchi Laboratory|Life Science and Technology News
- Researcher Profile|Tokyo Tech STAR Search - Hideki Taguchi
- Researcher Profile|Tokyo Tech STAR Search - Yuhei Chadani
- Researcher Profile|Tokyo Tech STAR Search - Tatsuya Niwa
- New insights into Protein Synthesis│Tokyo Tech News|Tokyo Tech News
- Institute of Innovative Research
- Department of Life Science and Technology, School of Life Science and Technology
- Latest Research News
Further Information
Professor Hideki Taguchi
School of Life Science and Technology,
Tokyo Institute of Technology
Email taguchi@bio.titech.ac.jp
Tel +81-45-924-5785