Life Science and Technology News
Pioneering Discovery of an Odor-detecting Receptor Enhancer
Scientists at Tokyo Tech have identified a regulatory sequence that turns gene expression on, or simply an enhancer, for odor-detecting receptors, which form one of the largest gene clusters in the mouse genome. This was done using a combination of research methods, including the CRISPR-Cas9 system, which is a genome editing technique, the Bacillus subtilis synthetic genome vector system, which is a cloning system for large DNA fragments, and bioinformatics.
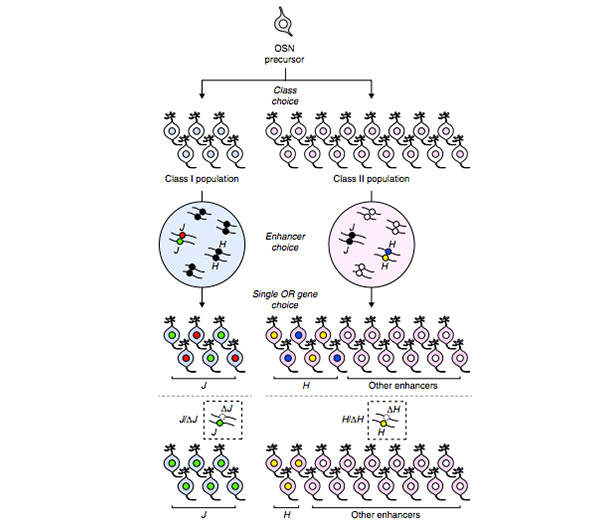
Figure 1. Mechanism by which J element determines expression of olfactory receptor genes.
Olfactory sensory neurons express either a class I or class II olfactory receptor gene in order to express either class I (blue) or class II (pink) olfactory sensory neurons. The enhancer is chosen based on the category, and class I sensory neurons activate one enhancer allele of the J element (green or red circle). On the other hand, class II sensory neurons activate other class II enhancers (white circle, H element is yellow or blue).
Each odor-detecting neuron (referred to as olfactory sensory neuron from here on), chooses a single odorant receptor gene from a fairly large number of options that are split into class I (fish-like) and class II (terrestrial-specific) odorant receptors. This strict selectiveness of sensory neurons is in part due to enhancers (DNA sequences that enhance transcription of a gene when bound by specific protein), which remain poorly understood. Understanding enhancer functions is of great interest due to their importance in gene expression as well as evolution and disease. However, they have not been sufficiently studied because they cannot be easily predicted from DNA sequences or chromosome modifications nor can they be easily identified.
The findings are especially important as they highlight the discovery of a regulatory sequence, termed the J element, that controls class I gene expression of many more genes than the counterparts that regulate class II gene expression. This extraordinary long-range regulation has never been seen before. The researchers also report that it is evolutionarily conserved across mammalian species, ranging from the most primitive mammal, the platypus, to humans. This may be especially useful as it sheds light onto why class I genes remain on a single region on one chromosome during mammalian evolution, while class II genes do not.
Based on genetic evidence, the findings also highlight the concept of allelic exclusion, or the expression of just one copy of a gene and not the other. This is especially important for the J element, as it shows that its activity determines which copy of a gene is expressed. This is a novel mechanism of expression of these receptor genes that is based on the gene regulatory region that determines if genes will be turned on or off.
The study "A long-range cis-regulatory element for class I odorant receptor genes" published in Nature Communications was a collaborative effort between Tokyo Tech, the University of Tokyo, Nihon BioData Corporation and RIKEN Brain Science Institute. Junji Hirota at Tokyo Tech and team focused on discovering a long-range enhancer for a large gene cluster, finding an evolutionary conserved sequence motif in mammalian evolution, and elucidating enhancer-dependent allelic preference or exclusion mechanism for odor-detecting receptor genes. Their findings point to a conserved sequence among mammalian genomes that was only present in the J element and not in any other class II elements. They speculate that the investigation of factors that bind this specific element could help in elucidating molecular mechanisms that drive the selective J-element dependent gene expression.
Reference
| Authors: | Tetsuo Iwata1, Yoshihito Niimura2,3, Chizuru Kobayashi4, Daichi Shirakawa4, Hikoyu Suzuki5, Takayuki Enomoto1, Kazushige Touhara2,3, Yoshihiro Yoshihara3,6 & Junji Hirota1,4,* |
|---|---|
| Title of original paper: | A long-range cis-regulatory element for class I odorant receptor genes |
| Journal: | Nature Communications |
| DOI: | 10.1038/s41467-017-00870-4 |
| Affiliations: |
1 Center for Biological Resources and Informatics, Tokyo Institute of Technology 2 Department of Applied Biological Chemistry, Graduate School of Agricultural and Life Sciences, The University of Tokyo 3 ERATO Touhara Chemosensory Signal Project, The Japan Science and Technology Agency, The University of Tokyo 4 Department of Life Science and Technology, Graduate School of Life Science and Technology, Tokyo Institute of Technology 5 Nihon BioData Corporation 6 RIKEN Brain Science Institute |
* Corresponding author's email:jhirota@bio.titech.ac.jp
- The Hirota Laboratory, Laboratory of Molecular Newroscience (Japanese)
- Researcher Profile | Tokyo Tech STAR Search - Junji Hirota
- Center for Biological Resources and Informatics
- Latest Research News
Further information
Associate Professor Junji Hirota
Center for Biological Resouces and Informatics,
Tokyo Institute of Technology
Email jhirota@bio.titech.ac.jp
Tel +81-45-924-5830