Life Science and Technology News
【Labs spotlight】 Nakamura and Yatsunami Laboratory
Extremophiles have limitless possibilities!
The Department has a variety of laboratories for Life Science and Technology, in which cutting-edge innovative research is being undertaken not only in basic science and engineering but also in the areas of medicine, pharmacy, agriculture, and multidisciplinary sciences.
This "Spotlight" series features a laboratory from the Department and introduces you to the laboratory's research projects and outcomes. This time we focus on Nakamura and Yatsunami Laboratory.
※Professor Nakamura was retired on March 31, 2020.

Life Science and Technology
Professor Satoshi Nakamura and Associate Professor Rie Yatsunami![]()
| Degree | Professor Satoshi Nakamura : PhD 1989, Tokyo Institute of Technology Associate Professor Rie Yatsunami : PhD 2000, Tokyo Institute of Technology |
|---|---|
| Areas of Research | Genetic engineering, Protein engineering, Directed evolution, Metabolic engineering |
| Keywords | Extremophiles, Extremozymes, Alkaliphiles, Halophiles, Thermophiles, Carotenoid biosynthtesis, Glycogen biosynthtesis |
| Website | Nakamura and Yatsunami Laboratory (Japanese) |
Research interest
We are interested in extremozymes and functional proteins produced by extremophiles, such as alkaliphiles, halophiles, thermophiles, using genetic engineering, protein engineering and directed evolution.
Extremozymes and functional proteins include xylanase, chitinase, chitosanase, beta-1,3-glucanase, alpha-amylase, glucoamylase, 3-isopropylmalate dehydrogenase, arabinofuranosidase, ferredoxin, transducer, FtsZ, cruxrhodopsin, cruxhalorhodopsin, etc.
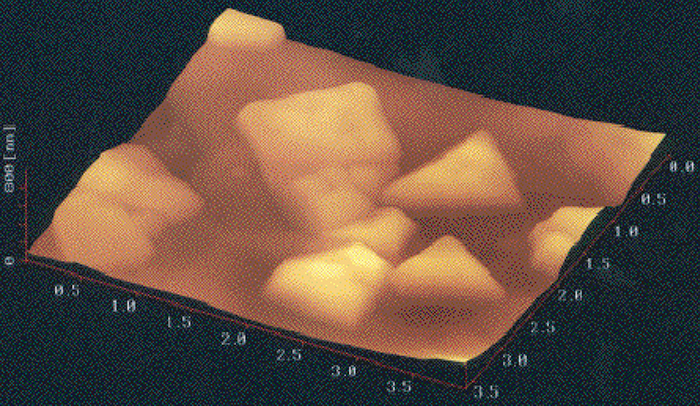
AFM image of a triangular disk-shaped extremely halophilic archaeon
Research findings
Selected publications
Satoshi Nakamura
- [1] Yang, Y., Yatsunami, R., Ando, A., Miyoko, N., Fukui, Takaichi, S., Nakamura, S. Complete biosynthetic pathway of the C50 carotenoid bacterioruberin from lycopene in the extremely halophilic archaeon Haloarcula japonica. J. Bacteriol., 197, 1614-1623 (2015).
- [2] Yatsunami, R., Ando, A., Yang, Y., Takaichi, S., Kohno, M., Matsumura, Y., Ikeda, H., Fukui, T., Nakasone, K., Fujita, N., Sekine, M., T. Takashina, T., Nakamura, S. Identification of carotenoids from the extremely halophilic archaeon Haloarcula japonica. Front. Microbiol., 5, Article 100 (2014).
- [3] Onodera, M., Yatsunami, R., Tsukimura, W., Fukui, T., Nakasone, K., Takashina, T., Nakamura, S. Gene analysis, expression, and characterization of an intracellular a-amylase from the extremely halophilic archaeon Haloarcula japonica. Biosci. Biotechnol. Biochem. 77, 281-288 (2013)
- [4] Uni, F., Lee, S., Yatsunami, R., Fukui, T., Nakamura, S. Mutational analysis of a CBM family 5 chitin-binding domain of an alkaline chitinase from Bacillus sp. J813. Biosci. Biotechnol. Biochem. 76, 530-535 (2012)
- [5] Yazawa, R., Takakura, J., Sakata, T., Ihsanawati, Yatsunami, R., Fukui, T., Kumasaka, T., Tanaka N., Nakamura, S. A calcium-dependent xylan-binding domain of alkaline xylanase from alkaliphilic Bacillus sp. strain 41M-1. Biosci. Bitechnol. Biochem. 75, 379-381 (2011)
- [6] Tsukimura, W., Watanabe, K., Morokuma, C., Yatsunami, R., Fukui T., Nakamura, S. Improvement of alkaliphily of thermostable GH family 10 xylanase from Thermotoga maritima by directed evolution, J. Jpn. Soc. Extr. 9, 15-18 (2010)
Rie Yatsunami
- [1] Endo, K., Nagao, K., Toyama, K., Ishida, S., Fukazawa, T., Yatsunami, R., Fukui, T., Xing, X., Nakamura, S. Cloning of a chitinase gene from newly isolated alkaliphilic Nocardiopsis sp. strain F96 and extracellular production of the recombinant protein in Escherichia coli. J. Jpn. Soc. Extremophiles, 14, 21-29 (2015).
- [2] Yang, Y., Yatsunami, R., Ando, A., Miyoko, N., Fukui, T., Takaichi, S. Nakamura, S. Complete Biosynthetic Pathway of the C50 Carotenoid Bacterioruberin from Lycopene in the Extremely Halophilic Archaeon Haloarcula japonica. J. Bacteriol., 197, 1614-1623 (2015).
- [3] Yatsunami, R., Ando, A., Yang, Y., Takaichi, S., Kohno, M., Matsumura, Y., Ikeda, H., Fukui, T., Nakasone, K., Fujita, N., Sekine, M., Takashina, T., Nakamura S. Identification of carotenoids from the extremely halophilic archaeon Haloarcula japonica. Front. Microbiol., 5, 100-105 (2014).
- [4] Tadikara, T., Matsubara, T., Kubota, Y., Kosaka, T., Ozawa, T., Tsukimura, W., Yatsunami, R., Fukui, T., Nakasone, K., Takashina, T., Nakamura, S. Gene expression and characterization of aerotaxis transducer HemAT from extremely halohilic archaeon Haloarcula japonica TR-1. J. Jpn. Soc. Extremophiles. 12, 29-32 (2013).
- [5] Onodera, M., Yatsunami, R., Tsukimura, W., Fukui, T., Nakasone, K., Takashina, T., Nakamura, S. Gene analysis, expression and characterization of an intracellular α-amylase from extremely halophilic archaeon Haloarcula japonica. Biosci. Biotechnol. Biochem., 77, 281-288 (2013).
- [6] Uni, F., Lee, S., Yatsuanmi, R., Fukui, T., Nakamura, S. Mutational Analysis of a CBM Family 5 Chitin-Binding Domain of an Alkaline Chitinase from Bacillus sp. J813. Biosci. Biotechnol. Biochem., 76, 530-535 (2012).
- [7] Chen, S., Ye, F., Chen Y., Zhao, H., Yatsunami, R., Nakamura, S., Arisaka, F., Xing, X.-H. Mechanism of the thermal inactivation of a MBP-fused heparinase I: Biochemical investigation and kinetic modeling. Biotechnol. Bioeng., 108, 1841-1851 (2011).
- [8] Yazawa, R., Takakura J., Sakata, T., Ihsanawati, Yatsunami, R., Fukui, T., Kumasaka,T., Tanaka, N. Nakamura, S. A Calcium-Dependent Xylan- Binding Domain of Alkaline Xylanase from Alkaliphilic Bacillus sp. Strain 41M-1. Biosci. Biotechnol. Biochem., 75, 379-381(2011).
Contact
Associate Professor Rie Yatsunami
Room 908, J2-J3 building, Suzukakedai campus
E-mail : yatsunami.r.aa@m.titech.ac.jp
*Find more about the lab and the latest activities at the lab site (Japanese)![]() .
.